The Python Book
All — By Topic
2019 2016 2015 2014
angle
argsort
beautifulsoup
binary
bisect
clean
collinear
covariance
cut_paste_cli
datafaking
dataframe
datetime
day_of_week
delta_time
df2sql
doctest
exif
floodfill
fold
format
frequency
gaussian
geocode
httpserver
is
join
legend
linalg
links
matrix
max
namedtuple
null
numpy
oo
osm
packaging
pandas
plot
point
range
regex
repeat
reverse
sample_words
shortcut
shorties
sort
stemming
strip_html
stripaccent
tools
visualization
zen
zip
3d aggregation angle archive argsort atan beautifulsoup binary bisect class clean collinear colsum comprehension count covariance csv cut_paste_cli datafaking dataframe datetime day_of_week delta_time deltatime df2sql distance doctest dotproduct dropnull exif file floodfill fold format formula frequency function garmin gaussian geocode geojson gps groupby html httpserver insert ipython is join kfold legend linalg links magic matrix max min namedtuple none null numpy onehot oo osm outer_product packaging pandas plot point quickies range read_data regex repeat reverse sample sample_data sample_words shortcut shorties sort split sqlite stack stemming string strip_html stripaccent tools track tuple visualization zen zip zipfile
Angle between 2 points
Calculate the angle in radians, between the horizontal through the 1st point and a line through the two points
def angle(p0,p1):
dx=float(p1.x)-float(p0.x)
dy=float(p1.y)-float(p0.y)
if dx==0:
if dy==0:
return 0.0
elif dy<0:
return math.atan(float('-inf'))
else:
return math.atan(float('inf'))
return math.atan(dy/dx)Get the indexes that would sort an array
Using numpy's argsort.
word_arr = np.array( ['lobated', 'demured', 'fristed', 'aproned', 'sheened', 'emulged',
'bestrid', 'mourned', 'upended', 'slashed'])
idx_sorted= np.argsort(word_arr)
idx_sorted
array([3, 6, 1, 5, 2, 0, 7, 4, 9, 8])Let's look at the first and last three elements:
print "First three :", word_arr[ idx_sorted[:3] ]
First three : ['aproned' 'bestrid' 'demured']
print "Last three :", word_arr[ idx_sorted[-3:] ]
Last three : ['sheened' 'slashed' 'upended']Index of min / max element
Using numpy's argmin.
Min:
In [4]: np.argmin(word_arr)
3
print word_arr[np.argmin(word_arr)]
apronedMax:
np.argmax(word_arr)
8
print word_arr[np.argmax(word_arr)]
upendedThe topics of the kubercon (Kubernetes conference)
Input
Markdown doc 'source.md' with all the presentation titles plus links:
[2000 Nodes and Beyond: How We Scaled Kubernetes to 60,000-Container Clusters
and Where We're Going Next - Marek Grabowski, Google Willow
A](/event/8K8w/2000-nodes-and-beyond-how-we-scaled-kubernetes-to-60000
-container-clusters-and-where-were-going-next-marek-grabowski-google) [How Box
Runs Containers in Production with Kubernetes - Sam Ghods, Box Grand Ballroom
D](/event/8K8u/how-box-runs-containers-in-production-with-kubernetes-sam-
ghods-box) [ITNW (If This Now What) - Orchestrating an Enterprise - Michael
Ward, Pearson Grand Ballroom C](/event/8K8t/itnw-if-this-now-what-
orchestrating-an-enterprise-michael-ward-pearson) [Unik: Unikernel Runtime for
Kubernetes - Idit Levine, EMC Redwood AB](/event/8K8v/unik-unikernel-runtime-
..
..Step 1: generate download script
Grab the links from 'source.md' and download them.
#!/usr/bin/python
# -*- coding: utf-8 -*-
import re
buf=""
infile = file('source.md', 'r')
for line in infile.readlines():
buf+=line.rstrip('\n')
oo=1
while True:
match = re.search( '^(.*?\()(/.[^\)]*)(\).*$)', buf)
if match is None:
break
url="https://cnkc16.sched.org"+match.group(2)
print "wget '{}' -O {:0>4d}.html".format(url,oo)
oo+=1
buf=match.group(3)Step 2: download the html
Execute the script generated by above code, and put the resulting files in directory 'content' :
wget 'https://cnkc16.sched.org/event/8K8w/2000-nodes-and-beyond-how-
we-scaled-kubernetes-to-60000-container-clusters-and-where-were-
going-next-marek-grabowski-google' -O 0001.html
wget 'https://cnkc16.sched.org/event/8K8u/how-box-runs-containers-in-
production-with-kubernetes-sam-ghods-box' -O 0002.html
wget 'https://cnkc16.sched.org/event/8K8t/itnw-if-this-now-what-
orchestrating-an-enterprise-michael-ward-pearson' -O 0003.html
.. Step 3: parse with beautiful soup
#!/usr/bin/python
# -*- coding: utf-8 -*-
from BeautifulSoup import *
import os
import re
import codecs
#outfile = file('text.md', 'w')
# ^^^ --> UnicodeEncodeError:
# 'ascii' codec can't encode character u'\u2019'
# in position 73: ordinal not in range(128)
outfile= codecs.open("text.md", "w", "utf-8")
file_ls=[]
for filename in os.listdir("content"):
if filename.endswith(".html"):
file_ls.append(filename)
for filename in sorted(file_ls):
infile = file('content/'+filename,'r')
content = infile.read()
infile.close()
soup = BeautifulSoup(content.decode('utf-8','ignore'))
div= soup.find('div', attrs={'class':'sched-container-inner'})
el_ls= div.findAll('span')
el=el_ls[0].text.strip()
title=re.sub(' - .*$','',el)
speaker=re.sub('^.* - ','',el)
outfile.write( u'\n\n## {}\n'.format(title))
outfile.write( u'\n\n{}\n'.format(speaker) )
det= div.find('div', attrs={'class':'tip-description'})
if det is not None:
outfile.write( u'\n{}\n'.format(det.text.strip() ) )Create a DB by scraping a webpage
Download all the webpages and put them in a zipfile (to avoid 're-downloading' on each try).
If you want to work 'direct', then use this to read the html content of a url:
html_doc=urllib.urlopen(url).read()Preparation: create database table
cur.execute('DROP TABLE IF EXISTS t_result')
cur.execute('''
CREATE TABLE t_result(
pos varchar(128),
nr varchar(128),
gesl varchar(128),
naam varchar(128),
leeftijd varchar(128),
ioc varchar(128),
tijd varchar(128),
tkm varchar(128),
gem varchar(128),
cat_plaats varchar(128),
cat_naam varchar(128),
gemeente varchar(128)
)
''') ## Pull each html file from the zipfile
zf=zipfile.ZipFile('brx_marathon_html.zip','r')
for fn in zf.namelist():
try:
content= zf.read(fn)
handle_content(content)
except KeyError:
print 'ERROR: %s not in zip file' % fn
breakParse the content of each html file with Beautiful Soup
soup = BeautifulSoup(content)
table= soup.find('table', attrs={'cellspacing':'0', 'cellpadding':'2'})
rows = table.findAll('tr')
for row in rows:
cols = row.findAll('td')
e = [ ele.text.strip() for ele in cols]
if len(e)>10:
cur.execute('INSERT INTO T_RESULT VALUES ( ?, ?, ?, ?, ?, ?, ?, ?, ?, ?, ?,? )',
(e[0],e[1],e[2],e[3],e[4],e[5],e[6],e[7],e[8],e[9],e[10],e[11]) )Note: the above code is beautiful soup 3, for beautiful soup 4, the findAll needs to be replaced by find_all.
Complete source code
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 | |
Binary vector
You have this vector that is a representation of a binary number. How to calculate the decimal value? Make the dot-product with the powers of two vector!
eg.
xbin=[1,1,1,1,1,0,1,0,0,0,0,0,0,0,0,0]
xdec=?Introduction:
import numpy as np
powers_of_two = (1 << np.arange(15, -1, -1))
array([32768, 16384, 8192, 4096, 2048, 1024, 512, 256, 128,
64, 32, 16, 8, 4, 2, 1])
seven=np.array( [0,0,0,0,0,0,0,0,0,0,0,0,0,1,1,1] )
seven.dot(powers_of_two)
7
thirtytwo=np.array( [0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0] )
thirtytwo.dot(powers_of_two)
32Solution:
xbin=np.array([1,1,1,1,1,0,1,0,0,0,0,0,0,0,0,0])
xdec=xbin.dot(powers_of_two)
=64000You can also write the binary vector with T/F:
xbin=np.array([True,True,True,True,True,False,True,False,
False,False,False,False,False,False,False,False])
xdec=xbin.dot(powers_of_two)
=64000Insert an element into an array, keeping the array ordered
Using the bisect_left() function of module bisect, which locates the insertion point.
def insert_ordered(ar, val):
i=0
if len(ar)>0:
i=bisect.bisect_left(ar,val)
ar.insert(i,val)Usage:
ar=[]
insert_ordered( ar, 10 )
insert_ordered( ar, 20 )
insert_ordered( ar, 5 )Retain recent files
eg. an application produces a backup file every night. You only want to keep the last 5 files.
Pipe the output of the following script (retain_recent.py) into a sh, eg in a cronjob:
/YOURPATH/retain_recent.py | bash Python code that produces 'rm fileX' statements:
#list the files according to a pattern, except for the N most recent files
import os,sys,time
fnList=[]
d='/../confluence_data/backups/'
for f in os.listdir( d ):
(mode, ino, dev, nlink, uid, gid, size, atime, mtime, ctime) = os.stat(d+"/"+f)
s='%s|%s/%s|%s' % ( mtime, d, f, time.ctime(mtime))
#print s
fnList.append(s)
# retain the 5 most recent files:
nd=len(fnList)-5
c=0
for s in sorted(fnList):
(mt,fn,hd)=s.split('|')
c+=1
if (c>=nd):
print '#keeping %s (%s)' % ( fn, hd )
else:
print 'rm -v %s #deleting (%s)' % ( fn, hd )Generate an array of collinear points plus some random points
- generate a number of points (integers) that are on the same line
- randomly intersperse these coordinates with a set of random points
- watchout: may generate dupes! (the random points, not the collinear points)
Source:
import random
p=[(5,5),(1,10)] # points that define the line
# warning: this won't work for vertical line!!!
slope= (float(p[1][1])-float(p[0][1]))/(float(p[1][0])-float(p[0][0]) )
intercept= float(p[0][1])-slope*float(p[0][0])
ar=[]
for x in range(0,25):
y=slope*float(x)+intercept
# only keep the y's that are integers
if (y%2)==0:
ar.append((x,int(y)))
# intersperse with random coordinates
r=3+random.randrange(0,5)
# only add random points when random nr is even
if r%2==0:
ar.extend( [ (random.randrange(0,100),random.randrange(0,100)) for j in range(r) ])
print arSample output:
[(1, 10), (97, 46), (94, 12), (33, 10), (9, 71), (9, 0), (28, 34),
(2, 94), (30, 29), (69, 28), (82, 31), (79, 86), (88, 46), (59, 24),
(2, 78), (54, 88), (94, 78), (99, 37), (75, 48), (91, 1), (67, 61),
(12, 11), (55, 55), (58, 82), (95, 99), (56, 27), (12, 18), (99, 25),
(77, 84), (31, 39), (64, 84), (4, 13), (80, 63), (43, 27), (78, 43),
(24, 32), (17, -10), (73, 15), (6, 97), (0, 74), (16, 97), (6, 77),
(60, 77), (19, 83), (19, 82), (19, 40), (58, 63), (64, 62), (14, 53),
(57, 21), (49, 24), (66, 94), (82, 1), (29, 39), (55, 64), (85, 68),
(39, 24)]np.random.multivariate_normal()
import numpy as np
import matplotlib.pyplot as plt
means = [
[9, 9], # top right
[1, 9], # top left
[1, 1], # bottom left
[9, 1], # bottom right
]
covariances = [
[ [.5, 0.], # covariance top right
[0, .5] ],
[[.1, .0], # covariance top left
[.0, .9]],
[[.9, 0.], # covariance bottom left
[0, .1]],
[[0.5, 0.5], # covariance bottom right
[0.5, 0.5]] ]
data = []
for k in range(len(means)):
for i in range(100) :
x = np.random.multivariate_normal(means[k], covariances[k])
data.append(x)
d=np.vstack(data)
plt.plot(d[:,0], d[:,1],'ko')
plt.show()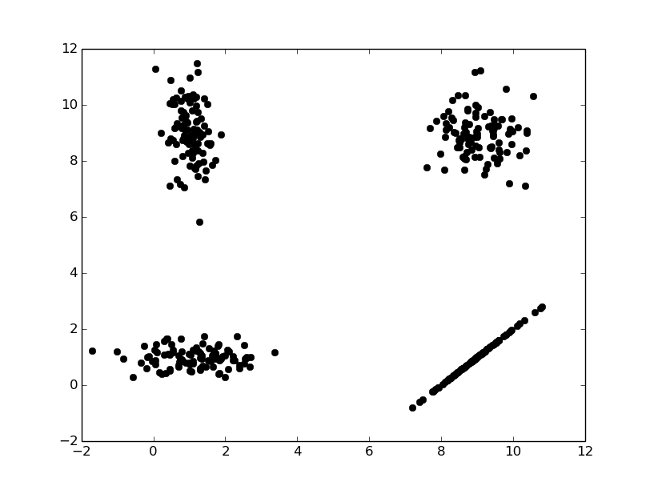
Cut and paste python on the command line
Simple example: number the lines in a text file
Read the file 'message.txt' and print a linenumber plus the line content.
python - message.txt <<EOF
import sys
i=1
with open(sys.argv[1],"r") as f:
for l in f.readlines():
print i,l.strip('\n')
i+=1
EOFOutput:
1 Better shutdown your ftp service.
2
3 W. Create a python program that reads a csv file, and uses the named fields
TBD.
Use namedtuple
Also see districtdatalabs.silvrback.com/simple-csv-data-wrangling-with-python
Anonymizing Data
Read this article on faker:
blog.districtdatalabs.com/a-practical-guide-to-anonymizing-datasets-with-python-faker
The goal: given a target dataset (for example, a CSV file with multiple columns), produce a new dataset such that for each row in the target, the anonymized dataset does not contain any personally identifying information. The anonymized dataset should have the same amount of data and maintain its analytical value. As shown in the figure below, one possible transformation simply maps original information to fake and therefore anonymous information but maintains the same overall structure.
Turn a dataframe into an array
eg. dataframe cn
cn
asciiname population elevation
128677 Crossett 5507 58
7990 Santa Maria da Vitoria 23488 438
25484 Shanling 0 628
95882 Colonia Santa Teresa 36845 2286
38943 Blomberg 1498 4
7409 Missao Velha 13106 364
36937 Goerzig 1295 81Turn into an arrary
cn.iloc[range(len(cn))].values
array([['Yuhu', 0, 15],
['Traventhal', 551, 42],
['Velabisht', 0, 60],
['Almorox', 2319, 539],
['Abuyog', 15632, 6],
['Zhangshan', 0, 132],
['Llica de Vall', 0, 136],
['Capellania', 2252, 31],
['Mezocsat', 6519, 91],
['Vars', 1634, 52]], dtype=object)Sidenote: cn was pulled from city data: cn=df.sample(7)[['asciiname','population','elevation']].
Filter a dataframe to retain rows with non-null values
Eg. you want only the data where the 'population' column has non-null values.
In short
df=df[df['population'].notnull()]Alternative, replace the null value with something:
df['population']=df['population'].fillna(0) In detail
import numpy as np
import pandas as pd
# setup the dataframe
data=[[ 'Isle of Skye', 9232, 124 ],
[ 'Vieux-Charmont', np.nan, 320 ],
[ 'Indian Head', 3844, 35 ],
[ 'Cihua', np.nan, 178 ],
[ 'Miasteczko Slaskie', 7327, 301 ],
[ 'Wawa', np.nan, 7 ],
[ 'Bat Khela', 46079, 673 ]]
df=pd.DataFrame(data, columns=['asciiname','population','elevation'])
#display the dataframe
df
asciiname population elevation
0 Isle of Skye 9232.0 124
1 Vieux-Charmont NaN 320
2 Indian Head 3844.0 35
3 Cihua NaN 178
4 Miasteczko Slaskie 7327.0 301
5 Wawa NaN 7
# retain only the rows where population has a non-null value
df=df[df['population'].notnull()]
asciiname population elevation
0 Isle of Skye 9232.0 124
2 Indian Head 3844.0 35
4 Miasteczko Slaskie 7327.0 301
6 Bat Khela 46079.0 673Add records to a dataframe in a for loop
The easiest way to get csv data into a dataframe is:
pd.read_csv('in.csv')But how to do it if you need to massage the data a bit, or your input data is not comma separated ?
cols= ['entrydate','sorttag','artist','album','doi','tag' ]
df=pd.DataFrame( columns= cols )
for ..:
data = .. a row of data-fields separated by |
with each field still to be stripped
of leading & trailing spaces
df.loc[len(df)]=map(str.strip,data.split('|'))Dataframe quickies
Count the number of different values in a column of a dataframe
pd.value_counts(df.Age)Drop a column
df['Unnamed: 0'].head() # first check if it is the right one
del df['Unnamed: 0'] # drop itDays between dates
Q: how many days are there in between these days ?
'29 sep 2016', '7 jul 2016', '28 apr 2016', '10 mar 2016', '14 jan 2016'Solution:
from datetime import datetime,timedelta
a=map(lambda x: datetime.strptime(x,'%d %b %Y'),
['29 sep 2016', '7 jul 2016', '28 apr 2016', '10 mar 2016', '14 jan 2016'] )
def dr(ar):
if len(ar)>1:
print "{:%d %b %Y} .. {} .. {:%d %b %Y} ".format(
ar[0], (ar[0]-ar[1]).days, ar[1])
dr(ar[1:]) Output:
dr(a)
29 Sep 2016 .. 84 .. 07 Jul 2016
07 Jul 2016 .. 70 .. 28 Apr 2016
28 Apr 2016 .. 49 .. 10 Mar 2016
10 Mar 2016 .. 56 .. 14 Jan 2016 Dataframe with date-time index
Create a dataframe df with a datetime index and some random values: (note: see 'simpler' dataframe creation further down)
|
Output:
In [4]: df.head(10)
Out[4]:
value
2009-12-01 71
2009-12-02 92
2009-12-03 64
2009-12-04 55
2009-12-05 99
2009-12-06 51
2009-12-07 68
2009-12-08 64
2009-12-09 90
2009-12-10 57
[10 rows x 1 columns]Now select a week of data
|
Output: watchout selects 8 days!!
In [235]: df[d1:d2]
Out[235]:
value
2009-12-10 99
2009-12-11 70
2009-12-12 83
2009-12-13 90
2009-12-14 60
2009-12-15 64
2009-12-16 59
2009-12-17 97
[8 rows x 1 columns]
In [236]: df[d1:d1+dt.timedelta(days=7)]
Out[236]:
value
2009-12-10 99
2009-12-11 70
2009-12-12 83
2009-12-13 90
2009-12-14 60
2009-12-15 64
2009-12-16 59
2009-12-17 97
[8 rows x 1 columns]
In [237]: df[d1:d1+dt.timedelta(weeks=1)]
Out[237]:
value
2009-12-10 99
2009-12-11 70
2009-12-12 83
2009-12-13 90
2009-12-14 60
2009-12-15 64
2009-12-16 59
2009-12-17 97
[8 rows x 1 columns]Postscriptum: a simpler way of creating the dataframe
An index of a range of dates can also be created like this with pandas:
pd.date_range('20091201', periods=31)Hence the dataframe:
df=pd.DataFrame(np.random.randint(50,100,31), index=pd.date_range('20091201', periods=31))Deduce the year from day_of_week
Suppose we know: it happened on Monday 17 November. Question: what year was it?
import datetime as dt
for i in [ dt.datetime(yr,11,17) for yr in range(1970,2014)]:
if i.weekday()==0: print i
1975-11-17 00:00:00
1980-11-17 00:00:00
1986-11-17 00:00:00
1997-11-17 00:00:00
2003-11-17 00:00:00
2008-11-17 00:00:00Or suppose we want to know all mondays of November for the same year range:
for i in [ dt.datetime(yr,11,1) + dt.timedelta(days=dy)
for yr in range(1970,2014) for dy in range(1,30)] :
if i.weekday()==0: print i
1970-11-02 00:00:00
1970-11-09 00:00:00
1970-11-16 00:00:00
..
..Add/subtract a delta time
Problem
A number of photo files were tagged as follows, with the date and the time:
20151205_17h48-img_0098.jpg
20151205_18h20-img_0099.jpg
20151205_18h21-img_0100.jpg..
Turns out that they should be all an hour earlier (reminder: mixing pics from two camera's), so let's create a script to rename these files...
Solution
1. Start
Let's use pandas:
import datetime as dt
import pandas as pd
import re
df0=pd.io.parsers.read_table( '/u01/work/20151205_gran_canaria/fl.txt',sep=",", \
header=None, names= ["fn"])
df=df0[df0['fn'].apply( lambda a: 'img_0' in a )] # filter out certain pics 2. Make parseable
Now add a column to the dataframe that only contains the numbers of the date, so it can be parsed:
df['rawdt']=df['fn'].apply( lambda a: re.sub('-.*.jpg','',a))\
.apply( lambda a: re.sub('[_h]','',a))Result:
df.head()
fn rawdt
0 20151202_07h17-img_0001.jpg 201512020717
1 20151202_07h17-img_0002.jpg 201512020717
2 20151202_07h17-img_0003.jpg 201512020717
3 20151202_15h29-img_0004.jpg 201512021529
28 20151202_17h59-img_0005.jpg 2015120217593. Convert to datetime, and subtract delta time
Convert the raw-date to a real date, and subtract an hour:
df['adjdt']=pd.to_datetime( df['rawdt'], format('%Y%m%d%H%M'))-dt.timedelta(hours=1)Note 20190105: apparently you can drop the 'format' string:
df['adjdt']=pd.to_datetime( df['rawdt'])-dt.timedelta(hours=1) Result:
fn rawdt adjdt
0 20151202_07h17-img_0001.jpg 201512020717 2015-12-02 06:17:00
1 20151202_07h17-img_0002.jpg 201512020717 2015-12-02 06:17:00
2 20151202_07h17-img_0003.jpg 201512020717 2015-12-02 06:17:00
3 20151202_15h29-img_0004.jpg 201512021529 2015-12-02 14:29:00
28 20151202_17h59-img_0005.jpg 201512021759 2015-12-02 16:59:004. Convert adjusted date to string
df['adj']=df['adjdt'].apply(lambda a: dt.datetime.strftime(a, "%Y%m%d_%Hh%M") )We also need the 'stem' of the filename:
df['stem']=df['fn'].apply(lambda a: re.sub('^.*-','',a) )Result:
df.head()
fn rawdt adjdt \
0 20151202_07h17-img_0001.jpg 201512020717 2015-12-02 06:17:00
1 20151202_07h17-img_0002.jpg 201512020717 2015-12-02 06:17:00
2 20151202_07h17-img_0003.jpg 201512020717 2015-12-02 06:17:00
3 20151202_15h29-img_0004.jpg 201512021529 2015-12-02 14:29:00
28 20151202_17h59-img_0005.jpg 201512021759 2015-12-02 16:59:00
adj stem
0 20151202_06h17 img_0001.jpg
1 20151202_06h17 img_0002.jpg
2 20151202_06h17 img_0003.jpg
3 20151202_14h29 img_0004.jpg
28 20151202_16h59 img_0005.jpg 5. Cleanup
Drop columns that are no longer useful:
df=df.drop(['rawdt','adjdt'], axis=1)Result:
df.head()
fn adj stem
0 20151202_07h17-img_0001.jpg 20151202_06h17 img_0001.jpg
1 20151202_07h17-img_0002.jpg 20151202_06h17 img_0002.jpg
2 20151202_07h17-img_0003.jpg 20151202_06h17 img_0003.jpg
3 20151202_15h29-img_0004.jpg 20151202_14h29 img_0004.jpg
28 20151202_17h59-img_0005.jpg 20151202_16h59 img_0005.jpg6. Generate scripts
Generate the 'rename' script:
sh=df.apply( lambda a: 'mv {} {}-{}'.format( a[0],a[1],a[2]), axis=1)
sh.to_csv('rename.sh',header=False, index=False )Also generate the 'rollback' script (in case we have to rollback the renaming) :
sh=df.apply( lambda a: 'mv {}-{} {}'.format( a[1],a[2],a[0]), axis=1)
sh.to_csv('rollback.sh',header=False, index=False )First lines of the rename script:
mv 20151202_07h17-img_0001.jpg 20151202_06h17-img_0001.jpg
mv 20151202_07h17-img_0002.jpg 20151202_06h17-img_0002.jpg
mv 20151202_07h17-img_0003.jpg 20151202_06h17-img_0003.jpg
mv 20151202_15h29-img_0004.jpg 20151202_14h29-img_0004.jpg
mv 20151202_17h59-img_0005.jpg 20151202_16h59-img_0005.jpgTurn a dataframe into sql statements
The easiest way is to go via sqlite!
eg. the two dataframes udf and tdf.
import sqlite3
con=sqlite3.connect('txdb.sqlite')
udf.to_sql(name='t_user', con=con, index=False)
tdf.to_sql(name='t_transaction', con=con, index=False)
con.close()Then on the command line:
sqlite3 txdb.sqlite .dump > create.sql This is the created create.sql script:
PRAGMA foreign_keys=OFF;
BEGIN TRANSACTION;
CREATE TABLE "t_user" (
"uid" INTEGER,
"name" TEXT
);
INSERT INTO "t_user" VALUES(9000,'Gerd Abrahamsson');
INSERT INTO "t_user" VALUES(9001,'Hanna Andersson');
INSERT INTO "t_user" VALUES(9002,'August Bergsten');
INSERT INTO "t_user" VALUES(9003,'Arvid Bohlin');
INSERT INTO "t_user" VALUES(9004,'Edvard Marklund');
INSERT INTO "t_user" VALUES(9005,'Ragnhild Brännström');
INSERT INTO "t_user" VALUES(9006,'Börje Wallin');
INSERT INTO "t_user" VALUES(9007,'Otto Byström');
INSERT INTO "t_user" VALUES(9008,'Elise Dahlström');
CREATE TABLE "t_transaction" (
"xid" INTEGER,
"uid" INTEGER,
"amount" INTEGER,
"date" TEXT
);
INSERT INTO "t_transaction" VALUES(5000,9008,498,'2016-02-21 06:28:49');
INSERT INTO "t_transaction" VALUES(5001,9003,268,'2016-01-17 13:37:38');
INSERT INTO "t_transaction" VALUES(5002,9003,621,'2016-02-24 15:36:53');
INSERT INTO "t_transaction" VALUES(5003,9007,-401,'2016-01-14 16:43:27');
INSERT INTO "t_transaction" VALUES(5004,9004,720,'2016-05-14 16:29:54');
INSERT INTO "t_transaction" VALUES(5005,9007,-492,'2016-02-24 23:58:57');
INSERT INTO "t_transaction" VALUES(5006,9002,-153,'2016-02-18 17:58:33');
INSERT INTO "t_transaction" VALUES(5007,9008,272,'2016-05-26 12:00:00');
INSERT INTO "t_transaction" VALUES(5008,9005,-250,'2016-02-24 23:14:52');
INSERT INTO "t_transaction" VALUES(5009,9008,82,'2016-04-20 18:33:25');
INSERT INTO "t_transaction" VALUES(5010,9006,549,'2016-02-16 14:37:25');
INSERT INTO "t_transaction" VALUES(5011,9008,-571,'2016-02-28 13:05:33');
INSERT INTO "t_transaction" VALUES(5012,9008,814,'2016-03-20 13:29:11');
INSERT INTO "t_transaction" VALUES(5013,9005,-114,'2016-02-06 14:55:10');
INSERT INTO "t_transaction" VALUES(5014,9005,819,'2016-01-18 10:50:20');
INSERT INTO "t_transaction" VALUES(5015,9001,-404,'2016-02-20 22:08:23');
INSERT INTO "t_transaction" VALUES(5016,9000,-95,'2016-05-09 10:26:05');
INSERT INTO "t_transaction" VALUES(5017,9003,428,'2016-03-27 15:30:47');
INSERT INTO "t_transaction" VALUES(5018,9002,-549,'2016-04-15 21:44:49');
INSERT INTO "t_transaction" VALUES(5019,9001,-462,'2016-03-09 20:32:35');
INSERT INTO "t_transaction" VALUES(5020,9004,-339,'2016-05-03 17:11:21');
COMMIT;The script doesn't create the indexes (because of Index='False'), so here are the statements:
CREATE INDEX "ix_t_user_uid" ON "t_user" ("uid");
CREATE INDEX "ix_t_transaction_xid" ON "t_transaction" ("xid");Or better: create primary keys on those tables!
Run a doctest on your python src file
.. first include unit tests in your docstrings:
eg. the file 'mydoctest.py'
#!/usr/bin/python3
def fact(n):
'''
Factorial.
>>> fact(6)
720
>>> fact(7)
5040
'''
return n*fact(n-1) if n>1 else nRun the doctests:
python3 -m doctest mydoctest.pyOr from within python:
>>> import doctest
>>> doctest.testfile("mydoctest.py")(hmm, doesn't work the way it should... import missing?)
List EXIF details of a photo
This program walks a directory, and lists selected exif data.
Install module exifread first.
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 | |
Flood fill
- Turn lines of text into a grid[row][column] (taking care to pad the lines to get a proper rectangle)
- Central data structure for the flood-fill is a stack
- If the randomly chosen point is blank, then fill it, and push the coordinates of its 4 neighbours onto the stack
- Handle the neighbouring points the same way
Src:
#!/usr/bin/python
import random
lines='''
+++++++ ++++++++++ ++++++++++++++++++++ +++++
+ + + + + + + +
+ + + + + + +++++
+ + + ++++ + +
+++++++ + + +
+ ++++ + +
+ + + +
+ + + +
+ + + +
++++++++++ + +
+ +
+ +
++++++++++++ + +
+ + + +
+ + + +
+ + + +
++++++++++++ ++++++++++++++
'''.split("\n")
# maximum number of columns and rows
colmax= max( [ len(line) for line in lines ] )
rowmax=len(lines)
padding=' ' * colmax
grid= [ list(lines[row]+padding)[0:colmax] for row in range(0,rowmax) ]
for l in grid: print( ''.join(l) ) # print the grid
print '-' * colmax # print a separating line
# creat a stack, and put a random coordinate on it
pointstack=[]
pointstack.append( ( random.randint(0,colmax), # col
random.randint(0,rowmax) ) ) # row
# floodfill
while len(pointstack)>0:
(col,row)=pointstack.pop()
if col>=0 and col<colmax and row>=0 and row<rowmax:
if grid[row][col]==' ':
grid[row][col]='O'
if col<(colmax-1): pointstack.append( (col+1,row))
if col>0: pointstack.append( (col-1,row))
if row<(rowmax-1): pointstack.append( (col,row+1) )
if row>0: pointstack.append( (col,row-1) )
for l in grid: print( ''.join(l) ) # print the gridOutput of a few runs:
+++++++ ++++++++++ ++++++++++++++++++++ +++++
+ + + + +OOOOOOOOOOOOOOOOOO+ + +
+ + + + +OOOOOOOOOOOOOOOOOO+ +++++
+ + + ++++ +OOOOOOOOOOOOOOOOOO+
+++++++ + +OOOOOOOOOOOOOOOOOO+
+ ++++ +OOOOOOOOOOOOOOOOOO+
+ + +OOOOOOOOOOOOOOOOOO+
+ + +OOOOOOOOOOOOOOOOOO+
+ + +OOOOOOOOOOOOOOOOOO+
++++++++++ +OOOOOOOOOOOOOOOOOO+
+OOOOOOOOOOOOOOOOOO+
+OOOOOOOOOOOOOOOOOO+
++++++++++++ +OOOOOOOOOOOOOOOOOO+
+ + +OOOOOOOOOOOOOOOOOO+
+ + +OOOOOOOOOOOOOOOO+
+ + +OOOOOOOOOOOOOO+
++++++++++++ ++++++++++++++ Completely flooded:
OOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOO
+++++++OOOOOOOO++++++++++OOOOOOOOOOOOOOOO++++++++++++++++++++OOOO+++++OOO
+ +OOOOOOOO+OOOOOOOO+OOOOOOOOOOOOOOOO+ +OOOO+ +OOO
+ +OOOOOOOO+OOOOOOOO+OOOOOOOOOOOOOOOO+ +OOOO+++++OOO
+ +OOOOOOOO+OOOOO++++OOOOOOOOOOOOOOOO+ +OOOOOOOOOOOO
+++++++OOOOOOOO+OOOOOOOOOOOOOOOOOOOOOOOOO+ +OOOOOOOOOOOO
OOOOOOOOOOOOOOO+OOOOO++++OOOOOOOOOOOOOOOO+ +OOOOOOOOOOOO
OOOOOOOOOOOOOOO+OOOOOOOO+OOOOOOOOOOOOOOOO+ +OOOOOOOOOOOO
OOOOOOOOOOOOOOO+OOOOOOOO+OOOOOOOOOOOOOOOO+ +OOOOOOOOOOOO
OOOOOOOOOOOOOOO+OOOOOOOO+OOOOOOOOOOOOOOOO+ +OOOOOOOOOOOO
OOOOOOOOOOOOOOO++++++++++OOOOOOOOOOOOOOOO+ +OOOOOOOOOOOO
OOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOO+ +OOOOOOOOOOOO
OOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOO+ +OOOOOOOOOOOO
OOOOO++++++++++++OOOOOOOOOOOOOOOOOOOOOOOO+ +OOOOOOOOOOOO
OOOOO+ +OOOOOOOOOOOOOOOOOOOOOOOO+ +OOOOOOOOOOOO
OOOOO+ +OOOOOOOOOOOOOOOOOOOOOOOOO+ +OOOOOOOOOOOOO
OOOOO+ +OOOOOOOOOOOOOOOOOOOOOOOOOO+ +OOOOOOOOOOOOOO
OOOOO++++++++++++OOOOOOOOOOOOOOOOOOOOOOOOOOO++++++++++++++OOOOOOOOOOOOOOO
OOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOOCheck the indexes on k-fold split
Suppose you split a list of n words into splits of k=5, what are the indexes of the splits?
Pseudo-code:
for i in 0..5:
start = n*i/k
end = n*(i+1)/kDouble check
Double check the above index formulas with words which have the same beginletter in a split (for easy validation).
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 | |
Output:
Split 0 of 5, length 7 : ['argot', 'along', 'addax', 'azans', 'aboil', 'aband', 'ayelp']
Split 1 of 5, length 7 : ['erred', 'ester', 'ekkas', 'entry', 'eldin', 'eruvs', 'ephas']
Split 2 of 5, length 7 : ['imino', 'islet', 'inurn', 'iller', 'idiom', 'izars', 'iring']
Split 3 of 5, length 7 : ['oches', 'outer', 'odist', 'orbit', 'ofays', 'outed', 'owned']
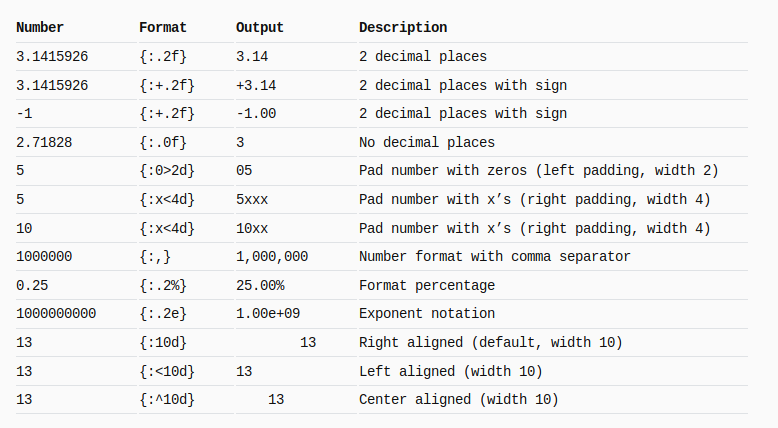
Split 4 of 5, length 7 : ['unlaw', 'upjet', 'upend', 'urged', 'urent', 'uncus', 'updry']Python string format
Following snap is taken from here: mkaz.blog/code/python-string-format-cookbook. Go there, there are more examples
Use the collections.counter to count the frequency of words in a text.
import collections
ln='''
The electrical and thermal conductivities of metals originate from
the fact that their outer electrons are delocalized. This situation
can be visualized by seeing the atomic structure of a metal as a
collection of atoms embedded in a sea of highly mobile electrons. The
electrical conductivity, as well as the electrons' contribution to
the heat capacity and heat conductivity of metals can be calculated
from the free electron model, which does not take into account the
detailed structure of the ion lattice.
When considering the electronic band structure and binding energy of
a metal, it is necessary to take into account the positive potential
caused by the specific arrangement of the ion cores - which is
periodic in crystals. The most important consequence of the periodic
potential is the formation of a small band gap at the boundary of the
Brillouin zone. Mathematically, the potential of the ion cores can be
treated by various models, the simplest being the nearly free
electron model.'''Split the text into words:
words=ln.lower().split()Create a Counter:
ctr=collections.Counter(words)Most frequent:
ctr.most_common(10)
[('the', 22),
('of', 12),
('a', 5),
('be', 3),
('by', 3),
('ion', 3),
('can', 3),
('and', 3),
('is', 3),
('as', 3)]Alternative: via df['col'].value_counts of pandas
import re
import pandas as pd
def removePunctuation(line):
return re.sub( "\s+"," ", re.sub( "[^a-zA-Z0-9 ]", "", line)).rstrip(' ').lstrip(' ').lower()
df=pd.DataFrame( [ removePunctuation(word.lower()) for word in ln.split() ], columns=['word'])
df['word'].value_counts()Result:
the 22
of 12
a 5
and 3
by 3
as 3
ion 3
..
..Plot a couple of Gaussians
import numpy as np
from math import pi
from math import sqrt
import matplotlib.pyplot as plt
def gaussian(x, mu, sig):
return 1./(sqrt(2.*pi)*sig)*np.exp(-np.power((x - mu)/sig, 2.)/2)
xv= map(lambda x: x/10.0, range(0,120,1))
mu= [ 2.0, 7.0, 9.0 ]
sig=[ 0.45, 0.70, 0.3 ]
for g in range(len(mu)):
m=mu[g]
s=sig[g]
yv=map( lambda x: gaussian(x,m,s), xv )
plt.plot(xv,yv)
plt.show()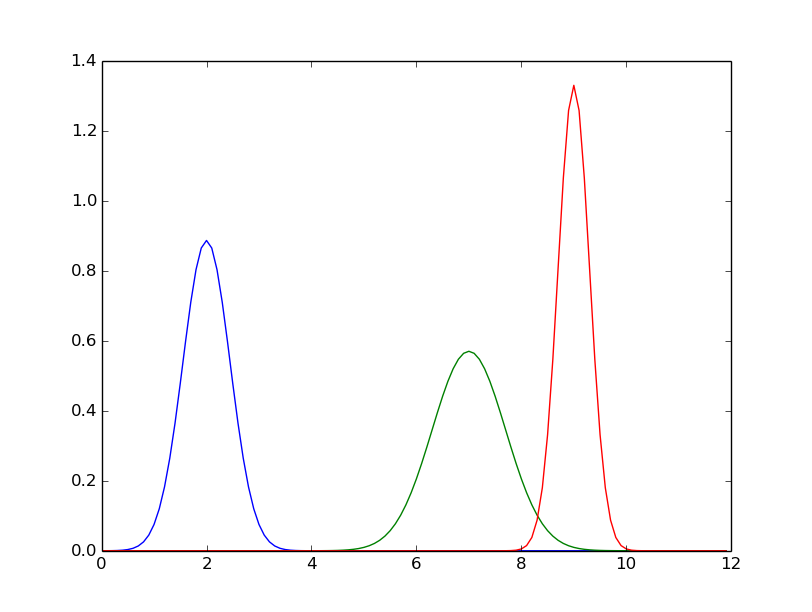
Incrementally update geocoded data
Startpoint: we have a sqlite3 database with (dirty) citynames and country codes. We would like to have the lat/lon coordinates, for each place.
Here some sample data, scraped from the Brussels marathon-results webpage, indicating the town of residence and nationality of the athlete:
LA CELLE SAINT CLOUD, FRA
TERTRE, BEL
FREDERICIA, DNK But sometimes the town and country don't match up, eg:
HEVERLEE CHN
WOLUWE-SAINT-PIERRE JPN
BUIZINGEN FRA For the geocoding we make use of nominatim.openstreetmap.org. The 3-letter country code also needs to be translated into a country name, for which we use the file /u01/data/20150215_country_code_iso/country_codes.txt.
The places for which we don't get valid (lat,lon) coordinates we put (0,0).
We run this script multiple times, small batches in the beginning, to be able see what the exceptions occur, and bigger batches in the end (when problems have been solved).
In between runs the data may be manually modified by opening the sqlite3 database, and updating/deleting the t_geocode table.
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 | |
A few queries
Total number of places:
select count(1) from t_geocode
916Number of places for which we didn't find valid coordinates :
select count(1) from t_geocode where lat=0 and lon=0
185Startup a simple http server
python -m SimpleHTTPServerAnd yes, that's all there is to it.
Only serves HEAD and GET, uses the current directory as root.
For python 3 it goes like this:
python3 -m http.server 5000is and '=='
From blog.lerner.co.il/why-you-should-almost-never-use-is-in-python
a="beta"
b=a
id(a)
3072868832L
id(b)
3072868832LIs the content of a and b the same? Yes.
a==b
TrueAre a and b pointing to the same object?
a is b
True
id(a)==id(b)
TrueSo it's safer to use '==', but for example for comparing to None, it's more readable and faster when writing :
if x is None:
print("x is None!")This works because the None object is a singleton.
Join two dataframes, sql style
You have a number of users, and a number of transactions against those users. Join these 2 dataframes.
import pandas as pd User dataframe
ids= [9000, 9001, 9002, 9003, 9004, 9005, 9006, 9007, 9008]
nms=[u'Gerd Abrahamsson', u'Hanna Andersson', u'August Bergsten',
u'Arvid Bohlin', u'Edvard Marklund', u'Ragnhild Br\xe4nnstr\xf6m',
u'B\xf6rje Wallin', u'Otto Bystr\xf6m',u'Elise Dahlstr\xf6m']
udf=pd.DataFrame(ids, columns=['uid'])
udf['name']=nmsContent of udf:
uid name
0 9000 Gerd Abrahamsson
1 9001 Hanna Andersson
2 9002 August Bergsten
3 9003 Arvid Bohlin
4 9004 Edvard Marklund
5 9005 Ragnhild Brännström
6 9006 Börje Wallin
7 9007 Otto Byström
8 9008 Elise DahlströmTransaction dataframe
tids= [5000, 5001, 5002, 5003, 5004, 5005, 5006, 5007, 5008, 5009, 5010, 5011, 5012,
5013, 5014, 5015, 5016, 5017, 5018, 5019, 5020]
uids= [9008, 9003, 9003, 9007, 9004, 9007, 9002, 9008, 9005, 9008, 9006, 9008, 9008,
9005, 9005, 9001, 9000, 9003, 9002, 9001, 9004]
tamt= [498, 268, 621, -401, 720, -492, -153, 272, -250, 82, 549, -571, 814, -114,
819, -404, -95, 428, -549, -462, -339]
tdt= ['2016-02-21 06:28:49', '2016-01-17 13:37:38', '2016-02-24 15:36:53',
'2016-01-14 16:43:27', '2016-05-14 16:29:54', '2016-02-24 23:58:57',
'2016-02-18 17:58:33', '2016-05-26 12:00:00', '2016-02-24 23:14:52',
'2016-04-20 18:33:25', '2016-02-16 14:37:25', '2016-02-28 13:05:33',
'2016-03-20 13:29:11', '2016-02-06 14:55:10', '2016-01-18 10:50:20',
'2016-02-20 22:08:23', '2016-05-09 10:26:05', '2016-03-27 15:30:47',
'2016-04-15 21:44:49', '2016-03-09 20:32:35', '2016-05-03 17:11:21']
tdf=pd.DataFrame(tids, columns=['xid'])
tdf['uid']=uids
tdf['amount']=tamt
tdf['date']=tdtContent of tdf:
xid uid amount date
0 5000 9008 498 2016-02-21 06:28:49
1 5001 9003 268 2016-01-17 13:37:38
2 5002 9003 621 2016-02-24 15:36:53
3 5003 9007 -401 2016-01-14 16:43:27
4 5004 9004 720 2016-05-14 16:29:54
5 5005 9007 -492 2016-02-24 23:58:57
6 5006 9002 -153 2016-02-18 17:58:33
7 5007 9008 272 2016-05-26 12:00:00
8 5008 9005 -250 2016-02-24 23:14:52
9 5009 9008 82 2016-04-20 18:33:25
10 5010 9006 549 2016-02-16 14:37:25
11 5011 9008 -571 2016-02-28 13:05:33
12 5012 9008 814 2016-03-20 13:29:11
13 5013 9005 -114 2016-02-06 14:55:10
14 5014 9005 819 2016-01-18 10:50:20
15 5015 9001 -404 2016-02-20 22:08:23
16 5016 9000 -95 2016-05-09 10:26:05
17 5017 9003 428 2016-03-27 15:30:47
18 5018 9002 -549 2016-04-15 21:44:49
19 5019 9001 -462 2016-03-09 20:32:35
20 5020 9004 -339 2016-05-03 17:11:21Join sql-style: pd.merge
pd.merge( tdf, udf, how='inner', left_on='uid', right_on='uid')
xid uid amount date name
0 5000 9008 498 2016-02-21 06:28:49 Elise Dahlström
1 5007 9008 272 2016-05-26 12:00:00 Elise Dahlström
2 5009 9008 82 2016-04-20 18:33:25 Elise Dahlström
3 5011 9008 -571 2016-02-28 13:05:33 Elise Dahlström
4 5012 9008 814 2016-03-20 13:29:11 Elise Dahlström
5 5001 9003 268 2016-01-17 13:37:38 Arvid Bohlin
6 5002 9003 621 2016-02-24 15:36:53 Arvid Bohlin
7 5017 9003 428 2016-03-27 15:30:47 Arvid Bohlin
8 5003 9007 -401 2016-01-14 16:43:27 Otto Byström
9 5005 9007 -492 2016-02-24 23:58:57 Otto Byström
10 5004 9004 720 2016-05-14 16:29:54 Edvard Marklund
11 5020 9004 -339 2016-05-03 17:11:21 Edvard Marklund
12 5006 9002 -153 2016-02-18 17:58:33 August Bergsten
13 5018 9002 -549 2016-04-15 21:44:49 August Bergsten
14 5008 9005 -250 2016-02-24 23:14:52 Ragnhild Brännström
15 5013 9005 -114 2016-02-06 14:55:10 Ragnhild Brännström
16 5014 9005 819 2016-01-18 10:50:20 Ragnhild Brännström
17 5010 9006 549 2016-02-16 14:37:25 Börje Wallin
18 5015 9001 -404 2016-02-20 22:08:23 Hanna Andersson
19 5019 9001 -462 2016-03-09 20:32:35 Hanna Andersson
20 5016 9000 -95 2016-05-09 10:26:05 Gerd AbrahamssonSidenote: fake data creation
This is the way the above fake data was created:
import random
from faker import Factory
fake = Factory.create('sv_SE')
ids=[]
nms=[]
for i in range(0,9):
ids.append(9000+i)
nms.append(fake.name())
print "%d\t%s" % ( ids[i],nms[i])
tids=[]
uids=[]
tamt=[]
tdt=[]
sign=[-1,1]
for i in range(0,21):
tids.append(5000+i)
tamt.append(sign[random.randint(0,1)]*random.randint(80,900))
uids.append(ids[random.randint(0,len(ids)-1)])
tdt.append(str(fake.date_time_this_year()))
print "%d\t%d\t%d\t%s" % ( tids[i], tamt[i], uids[i], tdt[i])Plot with simple legend
Use 'label' in your plot() call.
import math
import matplotlib.pyplot as plt
xv= map( lambda x: (x/4.)-10., range(0,81))
for l in [ 0.1, 0.5, 1., 5.] :
yv= map( lambda x: math.exp((-(-x)**2)/l), xv)
plt.plot(xv,yv,label='lambda = '+str(l));
plt.legend()
plt.show()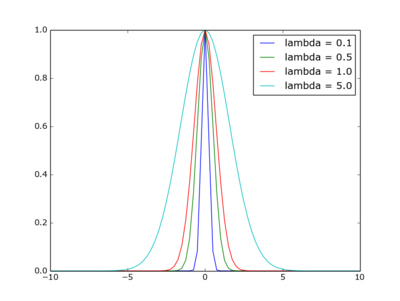
Sidenote: the function plotted is that of the Gaussian kernel in weighted nearest neighour regression, with xi=0
Linear Algebra MOOC
Odds of letters in scrable:
{'A':9/98, 'B':2/98, 'C':2/98, 'D':4/98, 'E':12/98, 'F':2/98,
'G':3/98, 'H':2/98, 'I':9/98, 'J':1/98, 'K':1/98, 'L':1/98,
'M':2/98, 'N':6/98, 'O':8/98, 'P':2/98, 'Q':1/98, 'R':6/98,
'S':4/98, 'T':6/98, 'U':4/98, 'V':2/98, 'W':2/98, 'X':1/98,
'Y':2/98, 'Z':1/98}Use // to find the remainder
Remainder using modulo: 2304811 % 47 -> 25
Remainder using // : 2304811 - 47 * (2304811 // 47) -> 25
Infinity
Infinity: float('infinity') : 1/float('infinity') -> 0.0
Set
Test membership with 'in' and 'not in'.
Note the difference between set (curly braces!) and tuple:
sum( {1,2,3,2} )
6
sum( (1,2,3,2) )
8Union of sets: use the bar operator { 1,2,4 } | { 1,3,5 } -> {1, 2, 3, 4, 5}
Intersection of sets: use the ampersand operator { 1,2,4 } & { 1,3,5 } -> {1}
Empty set: is not { } but set()! While for a list, the empty list is [].
Add / remove elements with .add() and .remove(). Add another set with .update()
s = { 1,2,4 }
s.update( { 1,3,5 } )
s
{1, 2, 3, 4, 5}Intersect with another set:
s.intersection_update( { 4,5,6,7 } )
s
{4, 5}Bind another variable to the same set: (any changes to s or t are visible to the other)
t=sMake a complete copy:
u=s.copy()Set comprehension:
{2*x for x in {1,2,3} }.. union of 2 sets combined with if (the if clause can be considered a filter) ..
s = { 1, 2,4 }
{ x for x in s|{5,6} if x>=2 }
{2, 4, 5, 6}Double comprehension : iterate over the Cartesian product of two sets:
{x*y for x in {1,2,3} for y in {2,3,4}}
{2, 3, 4, 6, 8, 9, 12}Compare to a list, which will return 2 more elements ( the 4 and the 6) :
[x*y for x in {1,2,3} for y in {2,3,4}]
[2, 3, 4, 4, 6, 8, 6, 9, 12]Or producing tuples:
{ (x,y) for x in {1,2,3} for y in {2,3,4}}
{(1, 2), (1, 3), (1, 4), (2, 2), (2, 3), (2, 4), (3, 2), (3, 3), (3, 4)}The factors of n:
n=64
{ (x,y) for x in range(1,1+int(math.sqrt(n))) for y in range(1,1+n) if (x*y)==n }
{(1, 64), (2, 32), (4, 16), (8, 8)}Or use it in a loop:
for n in range(40,100):
print (n ,
{ (x,y) for x in range(1,1+int(math.sqrt(n))) for y in range(1,1+n) if (x*y)==n })
40 {(4, 10), (1, 40), (2, 20), (5, 8)}
41 {(1, 41)}
42 {(1, 42), (2, 21), (6, 7), (3, 14)}
43 {(1, 43)}
44 {(2, 22), (1, 44), (4, 11)}
45 {(1, 45), (3, 15), (5, 9)}
46 {(2, 23), (1, 46)}
47 {(1, 47)}
48 {(3, 16), (2, 24), (1, 48), (4, 12), (6, 8)}
.. Lists
- a list can contain other sets and lists, but a set cannot contain a list (since lists are mutable).
- order: respected for lists, but not for sets
concatenate lists using the '+' operator
provide second argument
[]to sum to make this work:sum([ [1,2,3], [4,5,6], [7,8,9] ], [])->[1, 2, 3, 4, 5, 6, 7, 8, 9]
Skip elements in a slice: use the colon separate triple a:b:c notation
L = [0,10,20,30,40,50,60,70,80,90]
L[::3]
[0, 30, 60, 90]List of lists and comprehension:
listoflists = [[1,1],[2,4],[3, 9]]
[y for [x,y] in listoflists]
[1, 4, 9]Tuples
- difference with lists: a tuple is immutable. So sets may contain tuples.
Unpacking in a comprehension:
[y for (x,y) in [(1,'A'),(2,'B'),(3,'C')] ]
['A', 'B', 'C']
[x[1] for x in [(1,'A'),(2,'B'),(3,'C')] ]
['A', 'B', 'C']Converting list/set/tuple
Use constructors: set(), list() or tuple().
Note about range: a range represents a sequence, but it is not a list. Either iterate through the range or use set() or list() to turn it into a set or list.
Note about zip: it does not return a list, an 'iterator of tuples'.
Dictionary comprehensions
{ k:v for (k,v) in [(1,2),(3,4),(5,6)] }Iterate over the k,v pairs with items(), producing tuples:
[ item for item in {'a':1, 'b':2, 'c':3}.items() ]
[('a', 1), ('c', 3), ('b', 2)]Modules
- create your own module: name it properly, eg "spacerocket.py"
- import it in another script
While debugging it may be easier to use 'reload' (from package imp) to reload your module
Various
The enumerate function.
list(enumerate(['A','B','C']))
[(0, 'A'), (1, 'B'), (2, 'C')]
[ (i+1)*s for (i,s) in enumerate(['A','B','C','D','E'])]
['A', 'BB', 'CCC', 'DDDD', 'EEEEE']Python documentation links
- matplotlib
Add a column of zeros to a matrix
x= np.array([ [9.,4.,7.,3.], [ 2., 0., 3., 4.], [ 1.,2.,3.,1.] ])
array([[ 9., 4., 7., 3.],
[ 2., 0., 3., 4.],
[ 1., 2., 3., 1.]])Add the column:
np.c_[ np.zeros(3), x]
array([[ 0., 9., 4., 7., 3.],
[ 0., 2., 0., 3., 4.],
[ 0., 1., 2., 3., 1.]])Watchout: np.c_ takes SQUARE brackets, not parenthesis!
There is also an np.r_[ ... ] function. Maybe also have a look at vstack and hstack. See stackoverflow.com/a/8505658/4866785 for examples.
Matrix multiplication : dot product
a= np.array([[2., -1., 0.],[-3.,6.0,1.0]])
array([[ 2., -1., 0.],
[-3., 6., 1.]])
b= np.array([ [1.0,0.0,-1.0,2],[-4.,3.,1.,0.],[0.,3.,0.,-2.]])
array([[ 1., 0., -1., 2.],
[-4., 3., 1., 0.],
[ 0., 3., 0., -2.]])
np.dot(a,b)
array([[ 6., -3., -3., 4.],
[-27., 21., 9., -8.]])Dot product of two vectors
Take the first row of above a matrix and the first column of above b matrix:
np.dot( np.array([ 2., -1., 0.]), np.array([ 1.,-4.,0. ]) )
6.0Normalize a matrix
Normalize the columns: suppose the columns make up the features, and the rows the observations.
Calculate the 'normalizers':
norms=np.linalg.norm(a,axis=0)
print norms
[ 3.60555128 6.08276253 1. ]Turn a into normalized matrix an:
an = a/norms
print an
[[ 0.5547002 -0.16439899 0. ]
[-0.83205029 0.98639392 1. ]]Dot product used for aggregation of an unrolled matrix
Aggregations by column/row on an unrolled matrix, done via dot product. No need to reshape.
Column sums
Suppose this 'flat' array ..
a=np.array( [1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12 ] ).. represents an 'unrolled' 3x4 matrix ..
a.reshape(3,4)
array([[ 1, 2, 3, 4],
[ 5, 6, 7, 8],
[ 9, 10, 11, 12]]).. of which you want make the sums by column ..
a.reshape(3,4).sum(axis=0)
array([15, 18, 21, 24])This can also be done by the dot product of a tiled eye with the array!
np.tile(np.eye(4),3)
array([[ 1, 0, 0, 0, 1, 0, 0, 0, 1, 0, 0, 0],
[ 0, 1, 0, 0, 0, 1, 0, 0, 0, 1, 0, 0],
[ 0, 0, 1, 0, 0, 0, 1, 0, 0, 0, 1, 0],
[ 0, 0, 0, 1, 0, 0, 0, 1, 0, 0, 0, 1]])Dot product:
np.tile(np.eye(4),3).dot(a)
array([ 15., 18., 21., 24.])Row sums
Similar story :
a.reshape(3,4)
array([[ 1, 2, 3, 4],
[ 5, 6, 7, 8],
[ 9, 10, 11, 12]])Sum by row:
a.reshape(3,4).sum(axis=1)
array([10, 26, 42])Can be expressed by a Kronecker eye-onesie :
np.kron( np.eye(3), np.ones(4) )
array([[ 1, 1, 1, 1, 0, 0, 0, 0, 0, 0, 0, 0],
[ 0, 0, 0, 0, 1, 1, 1, 1, 0, 0, 0, 0],
[ 0, 0, 0, 0, 0, 0, 0, 0, 1, 1, 1, 1]])Dot product:
np.kron( np.eye(3), np.ones(4) ).dot(a)
array([ 10., 26., 42.])For the np.kron() function see Kronecker product
The dot product of two matrices (Eg. a matrix and it's tranpose), equals the sum of the outer products of the row-vectors & column-vectors.
a=np.matrix( "1 2; 3 4; 5 6" )
matrix([[1, 2],
[3, 4],
[5, 6]])Dot product of A and A^T :
np.dot( a, a.T)
matrix([[ 5, 11, 17],
[11, 25, 39],
[17, 39, 61]])Or as the sum of the outer products of the vectors:
np.outer(a[:,0],a.T[0,:])
array([[ 1, 3, 5],
[ 3, 9, 15],
[ 5, 15, 25]])
np.outer(a[:,1],a.T[1,:])
array([[ 4, 8, 12],
[ 8, 16, 24],
[12, 24, 36]]).. added up..
np.outer(a[:,0],a.T[0,:]) + np.outer(a[:,1],a.T[1,:])
array([[ 5, 11, 17],
[11, 25, 39],
[17, 39, 61]]).. and yes it is the same as the dot product!
Note: for above, because we are forming the dot product of a matrix with its transpose, we can also write it as (not using the transpose) :
np.outer(a[:,0],a[:,0]) + np.outer(a[:,1],a[:,1])Minimum and maximum int
- Max: sys.maxint
- Min: -sys.maxint-1
Named Tuple
Name the fields of your tuples
- namedtuple : factory function for creating tuple subclasses with named fields
- returns a new tuple subclass named typename.
- the new subclass is used to create tuple-like objects that have fields accessible by attribute lookup as well as being indexable and iterable.
Code:
import collections
Coord = collections.namedtuple('Coord', ['x','y'], verbose=False)
a=[ Coord(100.0,20.0), Coord(5.0,10.0), Coord(99.0,66.0) ]Access tuple elements by index:
a[1][0]
5.0Access tuple elements by name:
a[1].x
5.0Set verbose=True to see the code:
Coord = collections.namedtuple('Coord', ['x','y'], verbose=True)
class Coord(tuple):
'Coord(x, y)'
__slots__ = ()
_fields = ('x', 'y')
def __new__(_cls, x, y):
'Create new instance of Coord(x, y)'
return _tuple.__new__(_cls, (x, y))
@classmethod
def _make(cls, iterable, new=tuple.__new__, len=len):
'Make a new Coord object from a sequence or iterable'
result = new(cls, iterable)
if len(result) != 2:
raise TypeError('Expected 2 arguments, got %d' % len(result))
return result
def __repr__(self):
'Return a nicely formatted representation string'
return 'Coord(x=%r, y=%r)' % self
def _asdict(self):
'Return a new OrderedDict which maps field names to their values'
return OrderedDict(zip(self._fields, self))
def _replace(_self, **kwds):
'Return a new Coord object replacing specified fields with new values'
result = _self._make(map(kwds.pop, ('x', 'y'), _self))
if kwds:
raise ValueError('Got unexpected field names: %r' % kwds.keys())
return result
def __getnewargs__(self):
'Return self as a plain tuple. Used by copy and pickle.'
return tuple(self)
__dict__ = _property(_asdict)
def __getstate__(self):
'Exclude the OrderedDict from pickling'
pass
x = _property(_itemgetter(0), doc='Alias for field number 0')
y = _property(_itemgetter(1), doc='Alias for field number 1')'Null' value in Python is 'None'
There's always only one instance of this object, so you can check for equivalence with
x is None(identity comparison) instead ofx == None
stackoverflow.com/questions/3289601/null-object-in-python
Missing data in Python
Roughly speaking:
- missing 'object' -> None
- missing numerical value ->Nan ( np.nan )
Pandas handles both nearly interchangeably, converting if needed.
- isnull(), notnull() generates boolean mask indicating missing (or not) values
- dropna() return filtered version of data
- fillna() impute the missing values
More detail: jakevdp.github.io/PythonDataScienceHandbook/03.04-missing-values.html
Sample with replacement
Create a vector composed of randomly selected elements of a smaller vector. Ie. sample with replacement.
import numpy as np
src_v=np.array([1,2,3,5,8,13,21])
trg_v= src_v[np.random.randint( len(src_v), size=30)]
array([ 3, 8, 21, 5, 3, 3, 21, 5, 21, 3, 2, 13, 3, 21, 2, 2, 13,
5, 3, 21, 1, 2, 13, 3, 5, 3, 8, 8, 3, 1])Numpy quickies
Create a matrix of 6x2 filled with random integers:
import numpy as np
ra= np.matrix( np.reshape( np.random.randint(1,10,12), (6,2) ) )
matrix([[6, 1],
[3, 8],
[3, 9],
[4, 2],
[4, 7],
[3, 9]])The magic matrices (a la octave).
23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 | |
Sum column-wise (ie add up the elements for each column):
np.sum(magic3,axis=0)
array([15, 15, 15])Sum row-wise (ie add up elements for each row):
np.sum(magic3,axis=1)
array([15, 15, 15])Okay, a magic matrix is maybe not the best way to show row/column wise sums. Consider this:
rc= np.array([[0, 1, 2, 3, 4, 5],
[0, 1, 2, 3, 4, 5],
[0, 1, 2, 3, 4, 5]])
np.sum(rc,axis=0) # sum over rows
[0, 3, 6, 9, 12, 15]
np.sum(rc,axis=1) # sum over columns
[15,
15,
15]
np.sum(rc) # sum every element
45Simple OO program
Also from Dr. Chuck.
class PartyAnimal:
x = 0
def party(self) :
self.x = self.x + 1
print "So far",self.x
an = PartyAnimal()
an.party()
an.party()
an.party()Class with constructor / destructor
class PartyAnimal:
x = 0
def __init__(self):
print "I am constructed"
def party(self) :
self.x = self.x + 1
print "So far",self.x
def __del__(self):
print "I am destructed", self.x
an = PartyAnimal()
an.party()
an.party()
an.party()Field name added to Class
class PartyAnimal:
x = 0
name = ""
def __init__(self, nam):
self.name = nam
print self.name,"constructed"
def party(self) :
self.x = self.x + 1
print self.name,"party count",self.x
s = PartyAnimal("Sally")
s.party()
j = PartyAnimal("Jim")
j.party()
s.party()Inheritance
class PartyAnimal:
x = 0
name = ""
def __init__(self, nam):
self.name = nam
print self.name,"constructed"
def party(self) :
self.x = self.x + 1
print self.name,"party count",self.x
class FootballFan(PartyAnimal):
points = 0
def touchdown(self):
self.points = self.points + 7
self.party()
print self.name,"points",self.points
s = PartyAnimal("Sally")
s.party()
j = FootballFan("Jim")
j.party()
j.touchdown()Interesting blog post.
OpenStreetMap city blocks as GeoJSON polygons
Extracting blocks within a city as GeoJSON polygons from OpenStreetMap data
I'll talk about using QGIS software to explore and visualize LARGE maps and provide a Python script (you don't need QGIS for this) for converting lines that represent streets to polygons that represent city blocks. The script will use the polygonize function from Shapely but you need to preprocess the OSM data first which is the secret sauce.
peteris.rocks/blog/openstreetmap-city-blocks-as-geojson-polygons
Summary:
- Download GeoJSON files from Mapzen Metro Extracts
- Filter lines with filter.py
- Split LineStrings with multiple points to LineStrings with two points with split-lines.py
- Create polygons with polygonize.py
- Look at results with QGIS or geojson.io
GeoJSON: geojson.io/#map=14/-14.4439/28.4334
Intro:
I admit that I was a huge fan of the Python setuptools library for a long time. There was a lot in there which just resonated with how I thought that software development should work. I still think that the design of setuptools is amazing. Nobody would argue that setuptools was flawless and it certainly failed in many regards. The biggest problem probably was that it was build on Python's idiotic import system but there is only so little you can do about that. In general setuptools took the realistic approach to problem-solving: do the best you can do by writing a piece of software scratches the itch without involving a committee or require language changes. That also somewhat explains the second often cited problem of setuptools: that it's a monkeypatch on distutils.
Talks about: setuptools, distutils, .pth files, PIL, eggs, ..
Convert a Garmin .FIT file and plot the heartrate of your run
Use gpsbabel to turn your FIT file into CSV:
gpsbabel -t -i garmin_fit -f 97D54119.FIT -o unicsv -F 97D54119.csvPandas imports:
import pandas as pd
import math
import matplotlib.pyplot as plt
pd.options.display.width=150Read the CSV file:
df=pd.read_csv('97D54119.csv',sep=',',skip_blank_lines=False)Show some points:
df.head(500).tail(10)
No Latitude Longitude Altitude Speed Heartrate Cadence Date Time
490 491 50.855181 4.826737 78.2 2.79 144 78.0 2019/07/13 06:44:22
491 492 50.855136 4.826739 77.6 2.79 147 78.0 2019/07/13 06:44:24
492 493 50.854962 4.826829 76.2 2.77 148 77.0 2019/07/13 06:44:32
493 494 50.854778 4.826951 77.4 2.77 146 78.0 2019/07/13 06:44:41
494 495 50.854631 4.827062 78.0 2.71 143 78.0 2019/07/13 06:44:49
495 496 50.854531 4.827174 79.2 2.70 146 77.0 2019/07/13 06:44:54
496 497 50.854472 4.827249 79.2 2.73 149 77.0 2019/07/13 06:44:57
497 498 50.854315 4.827418 79.8 2.74 149 76.0 2019/07/13 06:45:05
498 499 50.854146 4.827516 77.4 2.67 147 76.0 2019/07/13 06:45:14
499 500 50.853985 4.827430 79.0 2.59 144 75.0 2019/07/13 06:45:22Function to compute the distance (approximately) :
# function to approximately calculate the distance between 2 points
# from: http://www.movable-type.co.uk/scripts/latlong.html
def rough_distance(lat1, lon1, lat2, lon2):
lat1 = lat1 * math.pi / 180.0
lon1 = lon1 * math.pi / 180.0
lat2 = lat2 * math.pi / 180.0
lon2 = lon2 * math.pi / 180.0
r = 6371.0 #// km
x = (lon2 - lon1) * math.cos((lat1+lat2)/2)
y = (lat2 - lat1)
d = math.sqrt(x*x+y*y) * r
return dCompute the distance:
ds=[]
(d,priorlat,priorlon)=(0.0, 0.0, 0.0)
for t in df[['Latitude','Longitude']].itertuples():
if len(ds)>0:
d+=rough_distance(t.Latitude,t.Longitude, priorlat, priorlon)
ds.append(d)
(priorlat,priorlon)=(t.Latitude,t.Longitude)
df['CumulativeDist']=ds Let's plot!
df.plot(kind='line',x='CumulativeDist',y='Heartrate',color='red')
plt.show() 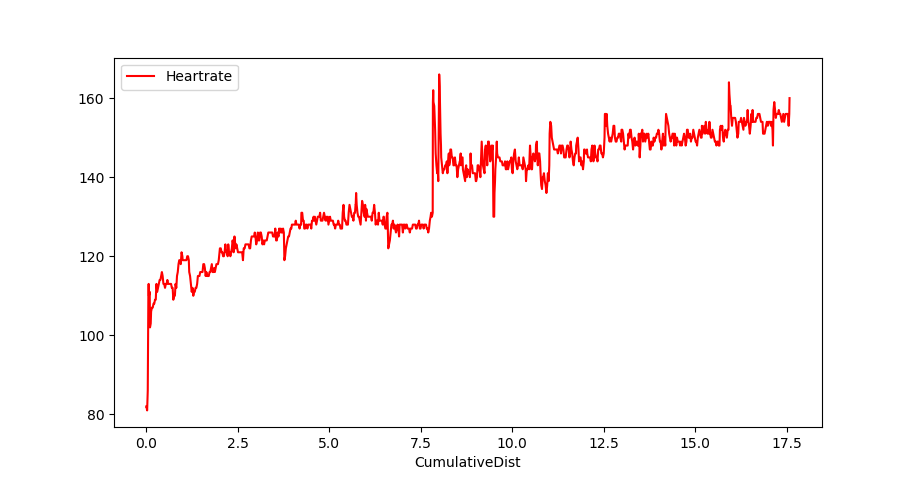
Or multiple columns:
plt.plot( df.CumulativeDist, df.Heartrate, color='red')
plt.plot( df.CumulativeDist, df.Altitude, color='blue')
plt.show()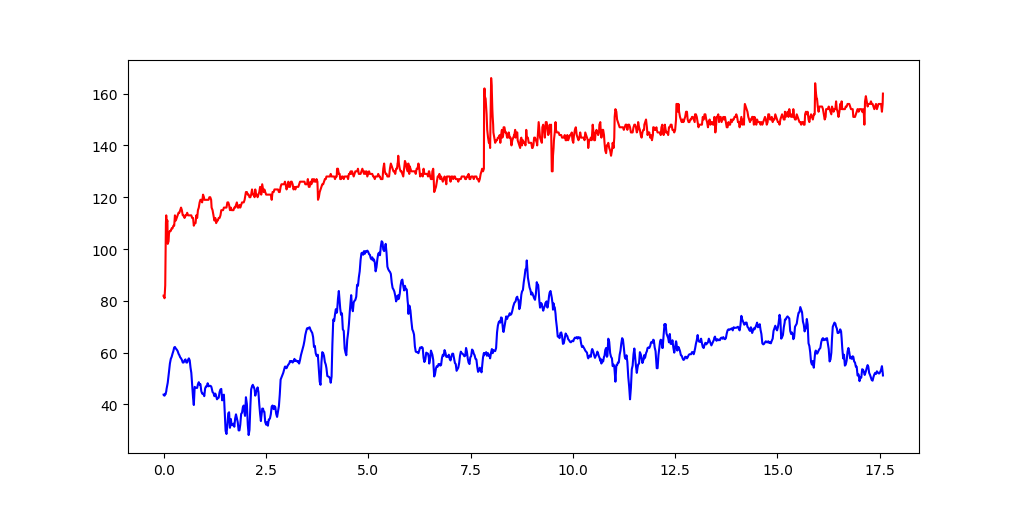
Turn a pandas dataframe into a dictionary
eg. create a mapping of a 3 digit code to a country name
BEL -> Belgium
CHN -> China
FRA -> France
..Code:
df=pd.io.parsers.read_table(
'/u01/data/20150215_country_code_iso/country_codes.txt',
sep='|')
c3d=df.set_index('c3')['country'].to_dict()Result:
c3d['AUS']
'Australia'
c3d['GBR']
'United Kingdom'eg. read a csv file that has nasty quotes, and save it as tab-separated.
import pandas as pd
import csv
colnames= ["userid", "movieid", "tag", "timestamp"]
df=pd.io.parsers.read_table("tags.csv",
sep=",", header=0, names= colnames,
quoting=csv.QUOTE_ALL)Write:
df.to_csv('tags.tsv', index=False, sep='\t')Dataframe aggregation fun
Load the city dataframe into dataframe df.
Summary statistic of 1 column
df.population.describe()
count 2.261100e+04
mean 1.113210e+05
std 4.337739e+05
min 0.000000e+00
25% 2.189950e+04
50% 3.545000e+04
75% 7.402450e+04
max 1.460851e+07Summary statistic per group
Load the city dataframe into df, then:
t1=df[['country','population']].groupby(['country'])
t2=t1.agg( ['min','mean','max','count'])
t2.sort_values(by=[ ('population','count') ],ascending=False).head(20)Output:
population
min mean max count
country
US 15002 62943.294138 8175133 2900
IN 15007 109181.708924 12691836 2398
BR 0 104364.320502 10021295 1195
DE 0 57970.979716 3426354 986
RU 15048 101571.065195 10381222 951
CN 15183 357967.030457 14608512 788
JP 15584 136453.906915 8336599 752
IT 895 49887.442136 2563241 674
GB 15024 81065.611200 7556900 625
FR 15009 44418.920455 2138551 616
ES 15006 65588.432282 3255944 539
MX 15074 153156.632735 12294193 501
PH 15066 100750.534884 10444527 430
TR 15058 142080.305263 11174257 380
ID 17504 170359.848901 8540121 364
PL 15002 64935.379421 1702139 311
PK 15048 160409.378641 11624219 309
NL 15071 53064.727626 777725 257
UA 15012 103468.816000 2514227 250
NG 15087 205090.336207 9000000 232Note on selecting a multilevel column
Eg. select 'min' via tuple ('population','min').
t2[ t2[('population','min')]>50000 ]
population
min mean max count
country
BB 98511 9.851100e+04 98511 1
CW 125000 1.250000e+05 125000 1
HK 288728 3.107000e+06 7012738 3
MO 520400 5.204000e+05 520400 1
MR 72337 3.668685e+05 661400 2
MV 103693 1.036930e+05 103693 1
SB 56298 5.629800e+04 56298 1
SG 3547809 3.547809e+06 3547809 1
ST 53300 5.330000e+04 53300 1
TL 150000 1.500000e+05 150000 1Read data from a zipfile into a dataframe
import pandas as pd
import zipfile
z = zipfile.ZipFile("lending-club-data.csv.zip")
df=pd.io.parsers.read_table(z.open("lending-club-data.csv"), sep=",")
z.close()Calculate the cumulative distance of gps trackpoints
Prep:
import pandas as pd
import mathFunction to calculate the distance:
# function to approximately calculate the distance between 2 points
# from: http://www.movable-type.co.uk/scripts/latlong.html
def rough_distance(lat1, lon1, lat2, lon2):
lat1 = lat1 * math.pi / 180.0
lon1 = lon1 * math.pi / 180.0
lat2 = lat2 * math.pi / 180.0
lon2 = lon2 * math.pi / 180.0
r = 6371.0 #// km
x = (lon2 - lon1) * math.cos((lat1+lat2)/2)
y = (lat2 - lat1)
d = math.sqrt(x*x+y*y) * r
return dRead data:
df=pd.io.parsers.read_table("trk.tsv",sep="\t")
# drop some columns (for clarity)
df=df.drop(['track','ele','tm_str'],axis=1) Sample:
df.head()
lat lon
0 50.848408 4.787456
1 50.848476 4.787367
2 50.848572 4.787275
3 50.848675 4.787207
4 50.848728 4.787189The prior-latitude column is the latitude column shifted by 1 unit:
df['prior_lat']= df['lat'].shift(1)
prior_lat_ix=df.columns.get_loc('prior_lat')
df.iloc[0,prior_lat_ix]= df.lat.iloc[0]The prior-longitude column is the longitude column shifted by 1 unit:
df['prior_lon']= df['lon'].shift(1)
prior_lon_ix=df.columns.get_loc('prior_lon')
df.iloc[0,prior_lon_ix]= df.lon.iloc[0]Calculate the distance:
df['dist']= df[ ['lat','lon','prior_lat','prior_lon'] ].apply(
lambda r : rough_distance ( r[0], r[1], r[2], r[3]) , axis=1)Calculate the cumulative distance
cum=0
cum_dist=[]
for d in df['dist']:
cum=cum+d
cum_dist.append(cum)
df['cum_dist']=cum_distSample:
df.head()
lat lon prior_lat prior_lon dist cum_dist
0 50.848408 4.787456 50.848408 4.787456 0.000000 0.000000
1 50.848476 4.787367 50.848408 4.787456 0.009831 0.009831
2 50.848572 4.787275 50.848476 4.787367 0.012435 0.022266
3 50.848675 4.787207 50.848572 4.787275 0.012399 0.034665
4 50.848728 4.787189 50.848675 4.787207 0.006067 0.040732
df.tail()
lat lon prior_lat prior_lon dist cum_dist
1012 50.847164 4.788163 50.846962 4.788238 0.023086 14.937470
1013 50.847267 4.788134 50.847164 4.788163 0.011634 14.949104
1014 50.847446 4.788057 50.847267 4.788134 0.020652 14.969756
1015 50.847630 4.787978 50.847446 4.788057 0.021097 14.990853
1016 50.847729 4.787932 50.847630 4.787978 0.011496 15.002349Onehot encode the categorical data of a data-frame
.. using the pandas get_dummies function.
Data:
import StringIO
import pandas as pd
data_strio=StringIO.StringIO('''category reason species
Decline Genuine 24
Improved Genuine 16
Improved Misclassified 85
Decline Misclassified 41
Decline Taxonomic 2
Improved Taxonomic 7
Decline Unclear 41
Improved Unclear 117''')
df=pd.read_fwf(data_strio)One hot encode 'category':
cat_oh= pd.get_dummies(df['category'])
cat_oh.columns= map( lambda x: "cat__"+x.lower(), cat_oh.columns.values)
cat_oh
cat__decline cat__improved
0 1 0
1 0 1
2 0 1
3 1 0
4 1 0
5 0 1
6 1 0
7 0 1Do the same for 'reason' :
reason_oh= pd.get_dummies(df['reason'])
reason_oh.columns= map( lambda x: "rsn__"+x.lower(), reason_oh.columns.values)Combine
Combine the columns into a new dataframe:
ohdf= pd.concat( [ cat_oh, reason_oh, df['species']], axis=1)Result:
ohdf
cat__decline cat__improved rsn__genuine rsn__misclassified \
0 1 0 1 0
1 0 1 1 0
2 0 1 0 1
3 1 0 0 1
4 1 0 0 0
5 0 1 0 0
6 1 0 0 0
7 0 1 0 0
rsn__taxonomic rsn__unclear species
0 0 0 24
1 0 0 16
2 0 0 85
3 0 0 41
4 1 0 2
5 1 0 7
6 0 1 41
7 0 1 117 Or if the 'drop' syntax on the dataframe is more convenient to you:
ohdf= pd.concat( [ cat_oh, reason_oh,
df.drop(['category','reason'], axis=1) ],
axis=1)Read a fixed-width datafile inline
import StringIO
import pandas as pd
data_strio=StringIO.StringIO('''category reason species
Decline Genuine 24
Improved Genuine 16
Improved Misclassified 85
Decline Misclassified 41
Decline Taxonomic 2
Improved Taxonomic 7
Decline Unclear 41
Improved Unclear 117''')Turn the string_IO into a dataframe:
df=pd.read_fwf(data_strio)Check the content:
df
category reason species
0 Decline Genuine 24
1 Improved Genuine 16
2 Improved Misclassified 85
3 Decline Misclassified 41
4 Decline Taxonomic 2
5 Improved Taxonomic 7
6 Decline Unclear 41
7 Improved Unclear 117The "5-number" summary
df.describe()
species
count 8.000000
mean 41.625000
std 40.177952
min 2.000000
25% 13.750000
50% 32.500000
75% 52.000000
max 117.000000Drop a column
df=df.drop('reason',axis=1) Result:
category species
0 Decline 24
1 Improved 16
2 Improved 85
3 Decline 41
4 Decline 2
5 Improved 7
6 Decline 41
7 Improved 117Create an empty dataframe
10 11 12 13 14 15 16 17 | |
Other way
24 25 | |
Quickies
You want to pandas to print more data on your wide terminal window?
pd.set_option('display.line_width', 200)You want to make the max column width larger?
pd.set_option('max_colwidth',80)Add two dataframes
Add the contents of two dataframes, having the same index
a=pd.DataFrame( np.random.randint(1,10,5), index=['a', 'b', 'c', 'd', 'e'], columns=['val'])
b=pd.DataFrame( np.random.randint(1,10,3), index=['b', 'c', 'e'],columns=['val'])
a
val
a 5
b 7
c 8
d 8
e 1
b
val
b 9
c 2
e 5
a+b
val
a NaN
b 16
c 10
d NaN
e 6
a.add(b,fill_value=0)
val
a 5
b 16
c 10
d 8
e 6Read/write csv
Read:
pd.read_csv('in.csv')Write:
<yourdataframe>.to_csv('out.csv',header=False, index=False ) Load a csv file
Load the following csv file. Difficulty: the date is spread over 3 fields.
2014, 8, 5, IBM, BUY, 50,
2014, 10, 9, IBM, SELL, 20 ,
2014, 9, 17, PG, BUY, 10,
2014, 8, 15, PG, SELL, 20 ,The way I implemented it:
10 11 12 13 14 15 16 17 18 19 20 21 22 | |
An alternative way,... it's better because the date is converted on reading, and the dataframe is indexed by the date.
|
Pie chart
Make a pie-chart of the top-10 number of cities per country in the file cities15000.txt
import pandas as pd
import matplotlib.pyplot as pltLoad city data, but only the country column.
colnames= [ "country" ]
df=pd.io.parsers.read_table("/usr/share/libtimezonemap/ui/cities15000.txt",
sep="\t", header=None, names= colnames,
usecols=[ 8 ])Get the counts:
cnts=df['country'].value_counts()
total_cities=cnts.sum()
22598Keep the top 10:
t10=cnts.order(ascending=False)[:10]
US 2900
IN 2398
BR 1195
DE 986
RU 951
CN 788
JP 752
IT 674
GB 625
FR 616What are the percentages ? (to display in the label)
pct=t10.map( lambda x: round((100.*x)/total_cities,2)).values
array([ 12.83, 10.61, 5.29, 4.36, 4.21, 3.49, 3.33, 2.98, 2.77, 2.73])Labels: country-name + percentage
labels=[ "{} ({}%)".format(cn,pc) for (cn,pc) in zip( t10.index.values, pct)]
['US (12.83%)', 'IN (10.61%)', 'BR (5.29%)', 'DE (4.36%)', 'RU (4.21%)', 'CN (3.49%)',
'JP (3.33%)', 'IT (2.98%)', 'GB (2.77%)', 'FR (2.73%)']Values:
values=t10.values
array([2900, 2398, 1195, 986, 951, 788, 752, 674, 625, 616])Plot
plt.style.use('ggplot')
plt.title('Number of Cities per Country\nIn file cities15000.txt')
plt.pie(values,labels=labels)
plt.show()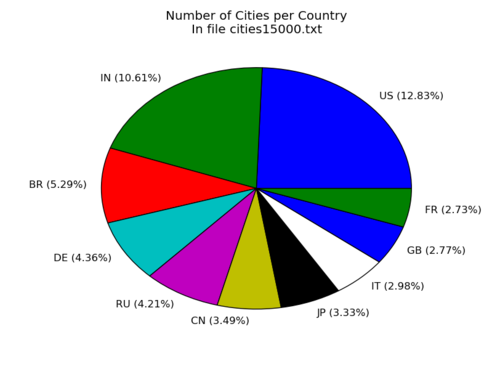
A good starting place:
matplotlib.org/mpl_toolkits/mplot3d/tutorial.html
Simple 3D scatter plot
Preliminary
from mpl_toolkits.mplot3d import axes3d
import matplotlib.pyplot as plt
import numpy as npData : create matrix X,Y,Z
X=[ [ i for i in range(0,10) ], ]*10
Y=np.transpose(X)
Z=[]
for i in range(len(X)):
R=[]
for j in range(len(Y)):
if i==j: R.append(2)
else: R.append(1)
Z.append(R)X:
[[0, 1, 2, 3, 4],
[0, 1, 2, 3, 4],
[0, 1, 2, 3, 4],
[0, 1, 2, 3, 4],
[0, 1, 2, 3, 4]]Y:
[[0, 0, 0, 0, 0],
[1, 1, 1, 1, 1],
[2, 2, 2, 2, 2],
[3, 3, 3, 3, 3],
[4, 4, 4, 4, 4]])Z:
[[2, 1, 1, 1, 1],
[1, 2, 1, 1, 1],
[1, 1, 2, 1, 1],
[1, 1, 1, 2, 1],
[1, 1, 1, 1, 2]]Scatter plot
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
ax.scatter(X, Y, Z)
plt.show()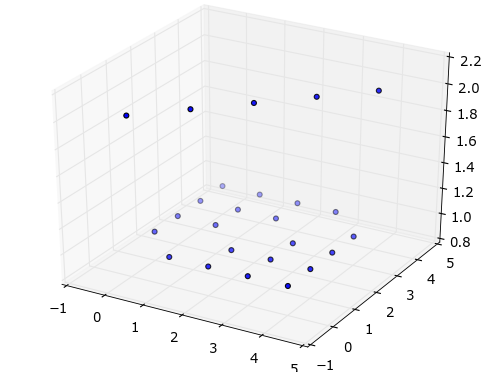
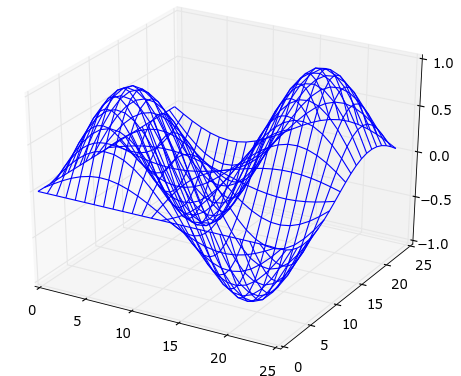
Wireframe plot
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 | |
Plot a function
eg. you want a plot of function: f(w) = 5-(w-10)² for w in the range 0..19
import matplotlib.pyplot as plt
x=range(20)
y=map( lambda w: 5-(w-10)**2, x)
plt.plot(x,y)
plt.show()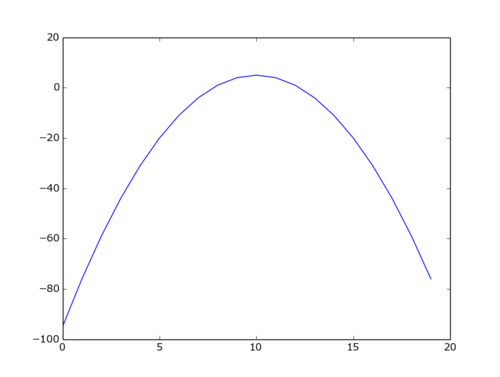
Plot some points
Imagine you have a list of tuples, and you want to plot these points:
l = [(1, 9), (2, 5), (3, 7)]And the plotting function expects to receive the x and y coordinate as separate lists.
First some fun with zip:
print(l)
[(1, 9), (2, 5), (3, 7)]
print(*l)
(1, 9) (2, 5) (3, 7)
print(*zip(*l))
(1, 2, 3) (9, 5, 7)Got it? Okay, let's plot.
plt.scatter(*zip(*pl))
plt.show()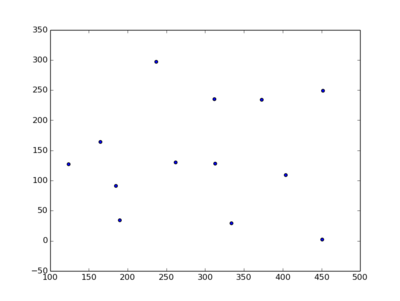
Define a point class
- with an x and y member
- with methods to 'autoprint'
Definition
import math
class P:
x=0
y=0
def __init__(self,x,y):
self.x=x
self.y=y
# gets called when a print is executed
def __str__(self):
return "x:{} y:{}".format(self.x,self.y)
# gets called eg. when a print is executed on an array of P's
def __repr__(self):
return "x:{} y:{}".format(self.x,self.y)
# convert an array of arrays or tuples to array of points
def convert(in_ar) :
out_ar=[]
for el in in_ar:
out_ar.append( P(el[0],el[1]) )
return out_arHow to initialize
Eg. create a list of points
# following initialisations lead to the same array of points (note the convert)
p=[P(0,0),P(0,1),P(-0.866025,-0.5),P(0.866025,0.5)]
q=convert( [[0,0],[0,1],[-0.866025,-0.5],[0.866025,0.5]] )
r=convert( [(0,0),(0,1),(-0.866025,-0.5),(0.866025,0.5)] )
print type(p), ' | ' , p[2].x, p[2].y, ' | ', p[2]
print type(q), ' | ' , q[2].x, q[2].y, ' | ', q[2]
print type(r), ' | ' , r[2].x, r[2].y, ' | ', r[2]Output:
<type 'list'> | -0.866025 -0.5 | x:-0.866025 y:-0.5
<type 'list'> | -0.866025 -0.5 | x:-0.866025 y:-0.5
<type 'list'> | -0.866025 -0.5 | x:-0.866025 y:-0.5How to use
eg. Calculate the angle between :
- the horizontal line through the first point
- and the line through the two points
Then convert the result from radians to degrees: (watchout: won't work for dx==0)
print math.atan( ( p[3].y - p[0].y ) / ( p[3].x - p[0].x ) ) * 180.0/math.piOutput:
30.0000115676Generate n numbers in an interval
Return evenly spaced numbers over a specified interval.
Pre-req:
import numpy as np
import matplotlib.pyplot as pltIn linear space
y=np.linspace(0,90,num=10)
array([ 0., 10., 20., 30., 40., 50., 60., 70., 80., 90.])
x=[ i for i in range(len(y)) ]
[0, 1, 2, 3, 4, 5, 6, 7, 8, 9]
plt.plot(x,y)
plt.scatter(x,y)
plt.title("linspace")
plt.show()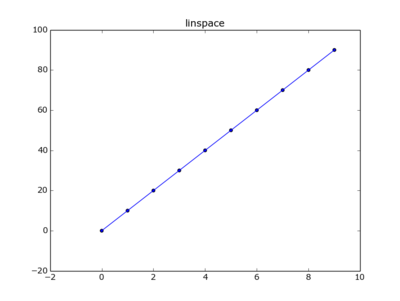
In log space
y=np.logspace(0, 9, num=10)
array([ 1.00000000e+00, 1.00000000e+01, 1.00000000e+02,
1.00000000e+03, 1.00000000e+04, 1.00000000e+05,
1.00000000e+06, 1.00000000e+07, 1.00000000e+08,
1.00000000e+09])
x=[ i for i in range(len(y)) ]
plt.plot(x,y)
plt.scatter(x,y)
plt.title("logspace")
plt.show()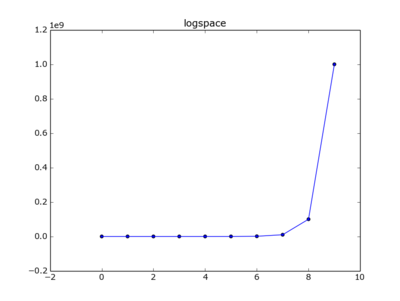
Plotting the latter on a log scale..
plt.plot(x,y)
plt.scatter(x,y)
plt.yscale('log')
plt.title("logspace on y-logscale")
plt.show()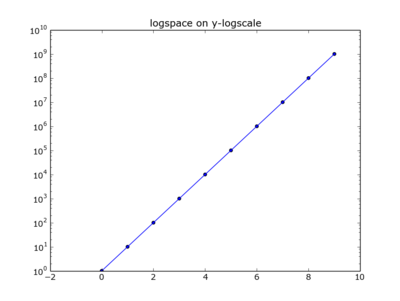
Here is Dr. Chuck's RegEx "cheat sheet". You can also download it here:
www.dr-chuck.net/pythonlearn/lectures/Py4Inf-11-Regex-Guide.doc

Here's Dr. Chucks book on learning python: www.pythonlearn.com/html-270
For more information about using regular expressions in Python, see docs.python.org/2/howto/regex.html
Regex: positive lookbehind assertion
(?<=...) Matches if the current position in the string is preceded by a match for ... that ends at the current position.
eg.
s="Yes, taters is a synonym for potaters or potatoes."
re.sub('(?<=po)taters','TATERS', s)
'Yes, taters is a synonym for poTATERS or potatoes.'Or example from python doc:
m = re.search('(?<=abc)def', 'abcdef')
m.group(0)
'def'Fill an array with 1 particular value
Via comprehension:
z1=[0 for x in range(20)]
z1
[0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0]Via built-in repeat:
z2=[0] * 20 Equal? yes.
z1==z2
TrueWhich one is faster?
timeit.timeit('z=[0 for x in range(100000)]',number=100)
0.8116392770316452
timeit.timeit('z=[0]*100000',number=100)
0.050275236018933356The built-in repeat beats comprehension hands down!
Reverse the words in a string
Tactic:
- reverse the whole string
- reverse every word on its own
Code:
s="The goal is to make a thin circle of dough, with a raised edge."
r=' '.join([ w[::-1] for w in s[::-1].split(' ') ]) The notation s[::-1] is called extended slice syntax, and should be read as [start:end:step] with the start and end left off.
Note: you also have the reversed() function, which returns an iterator:
s='circumference'
''.join(reversed(s))
'ecnerefmucric'Produce sample words
Use the sowpods file to generate a list of words that fulfills a special condition (eg length, starting letter) Use is made of the function random.sample(population, k) to take a unique sample of a larger list.
1 2 3 4 5 6 7 8 9 10 11 12 | |
Output:
['argot', 'along', 'addax', 'azans', 'aboil', 'aband']
['erred', 'ester', 'ekkas', 'entry', 'eldin', 'eruvs']
['imino', 'islet', 'inurn', 'iller', 'idiom', 'izars']
['oches', 'outer', 'odist', 'orbit', 'ofays', 'outed']
['unlaw', 'upjet', 'upend', 'urged', 'urent', 'uncus']Combinations of array elements
Suppose you want to know how many unique combinations of the elements of an array there are.
The scenic tour:
a=list('abcdefghijklmnopqrstuvwxyz')
n=len(a)
stack=[]
for i in xrange(n):
for j in xrange(i+1,n):
stack.append( (a[i],a[j]) )
print len(stack) The shortcut:
print (math.factorial(n)/2)/math.factorial(n-2)From the more general formula:
n! / r! / (n-r)! with r=2. (see itertools.combinations(iterable, r) on docs.python.org/2/library/itertools.html )
Add up from 1 to n
Add up all numbers from 1 to n.
The straight-n-simple solution:
result = 0
for i in xrange(n):
result += (i + 1)
print (result)The fast solution:
result = n * (n+1) / 2
print (result)Here's the explanation (thanks codility)
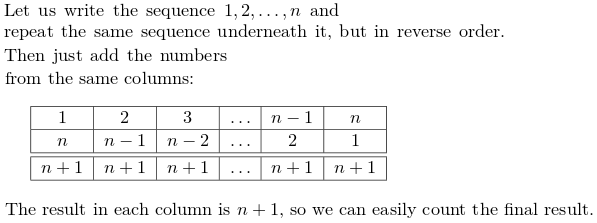

Shorties & quickies
Get an array of 10 random numbers
randrange: choose a random item from range(start, stop[)
import random
rand_arr=[ random.randrange(0,10) for i in range(10) ]Sort a list of tuples
Say you have markers that are tuples, and you want to have the marker list sorted by the first tuple element.
sorted_marker=sorted( marker, key=lambda x:x[0] ) Multi-level sort: custom compare
Use a custom compare function. The compare function receives 2 objects to be compared.
def cust_cmp(x,y):
if (x[1]==y[1]):
return cmp( x[0],y[0] )
return cmp(x[1],y[1])
names= [ ('mahalia', 'jackson'), ('moon', 'zappa'), ('janet','jackson'), ('lee','albert'), ('latoya','jackson') ]
sorted_names=sorted( names, cmp=cust_cmp)Output:
('lee', 'albert')
('janet', 'jackson')
('latoya', 'jackson')
('mahalia', 'jackson')
('moon', 'zappa')- stemmer that implements porter stemmer: stemming package
Strip HTML tags from a text.
6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 | |
Output:
The Köln concert
[edit]
The concert was organized by 17-year-old
Vera Brandes, then Germany ’s youngest concert promoter.
[5] At Jarrett's request, Brandes
had selected a Bösendorfer
290 Imperial concert grand piano for the performance. As found on : stackoverflow
Strip accents from letters
See how sklearn does it, functions:
strip_accents_ascii(s)
strip_accents_unicode(s)github.com/scikit-learn/scikit-learn/blob/master/sklearn/feature_extraction/text.py
See also: stackoverflow.com/questions/517923/what-is-the-best-way-to-remove-accents-in-a-python-unicode-string
Digest of infoworld article
- Beautiful Soup: Processing parse trees -- XML, HTML, or similarly structured data
- Pillow: image processin g(following to PIL)
- Gooey: turn a console-based Python program into one that sports a platform-native GUI.
- Peewee: a tiny ORM that supports SQLite, MySQL, and PostgreSQL, with many extensions.
- Scrapy: screen scraping and Web crawling.
- Apache Libcloud: accessing multiple cloud providers through a single, consistent, and unified API.
- Pygame: a framework for creating video games in Python.
- Pathlib: handling filesystem paths in a consistent and cross-platform way, courtesy of a module that is now an integral part of Python.
- NumPy: scientific computing and mathematical work, including statistics, linear algebra, matrix math, financial operations, and tons more.
- Sh: calling any external program, in a subprocess, and returning the results to a Python program -- but with the same syntax as if the program in question were a native Python function.
Visualizing distributions of data
Visualizing distributions of data
This notebook demonstrates different approaches to graphically representing distributions of data, specifically focusing on the tools provided by the seaborn packageb
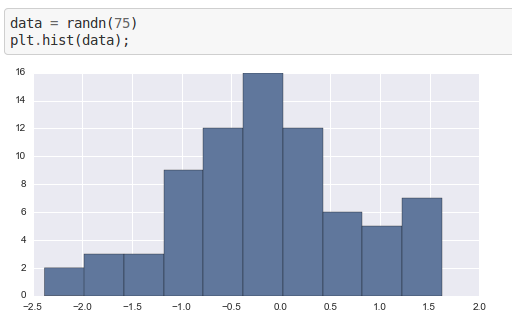
The Zen of Python
From: www.thezenofpython.com
Beautiful is better than ugly.
Explicit is better than implicit.
Simple is better than complex.
Complex is better than complicated.
Flat is better than nested.
Sparse is better than dense.
Readability counts.
Special cases aren't special enough to break the rules.
Although practicality beats purity.
Errors should never pass silently.
Unless explicitly silenced.
In the face of ambiguity, refuse the temptation to guess.
There should be one-- and preferably only one --obvious way to do it.
Although that way may not be obvious at first unless you're Dutch.
Now is better than never.
Although never is often better than *right* now.
If the implementation is hard to explain, it's a bad idea.
If the implementation is easy to explain, it may be a good idea.
Namespaces are one honking great idea -- let's do more of those!Archive files into a zipfile
A cronjob runs every 10 minutes produces these JSON files:
6752 Sep 10 08:30 orderbook-kraken-20150910-083003.json
6682 Sep 10 08:40 orderbook-kraken-20150910-084004.json
6717 Sep 10 08:50 orderbook-kraken-20150910-085004.json
6717 Sep 10 09:00 orderbook-kraken-20150910-090003.json
6682 Sep 10 09:10 orderbook-kraken-20150910-091003.json
6717 Sep 10 09:20 orderbook-kraken-20150910-092002.json
6717 Sep 10 09:30 orderbook-kraken-20150910-093003.json
6682 Sep 10 09:40 orderbook-kraken-20150910-094003.json
6682 Sep 10 09:50 orderbook-kraken-20150910-095003.json
6752 Sep 10 10:00 orderbook-kraken-20150910-100004.json
6788 Sep 10 10:10 orderbook-kraken-20150910-101003.json
6787 Sep 10 10:20 orderbook-kraken-20150910-102004.json
6823 Sep 10 10:30 orderbook-kraken-20150910-103004.json
6752 Sep 10 10:40 orderbook-kraken-20150910-104004.jsonAnother cronjob, run every morning, zips all the files of one day together into a zipfile, turning above files into:
20150910-orderbook-kraken.zip
20150911-orderbook-kraken.zip
20150912-orderbook-kraken.zip
.. Here's the code of the archiver (ie the 2nd cronjob) :
import os,re,sys
import zipfile
from datetime import datetime
# get a date from the filename. Assumptions:
# - format = YYYYMMDD
# - date is in this millenium (ie starts with 2)
# - first number in the filename is the date
def get_date(fn):
rv=re.sub(r'(^.[^0-9]*)(2[0-9]{7})(.*)', r'\2', fn)
return rv
# first of all set the working directory
wd=sys.argv[1]
if (len(wd)==0):
print "Need a working directory"
sys.exit(0)
os.chdir(wd)
# find the oldest date-pattern in the json files
ds=set()
for filename in os.listdir("."):
if filename.endswith(".json"):
ds.add(get_date(filename))
#print "{}->{}".format(filename,dt)
# exclude today's pattern (because today may not be complete):
today=datetime.now().strftime("%Y%m%d")
ds.remove(today)
l=sorted(list(ds))
#print l
if (len(l)==0):
#print "Nothing to do!"
sys.exit(0)
# datepattern selected
datepattern=l[0]
# decide on which files go into the archive
file_ls=[]
for filename in os.listdir("."):
if filename.endswith(".json") and filename.find(datepattern)>-1:
file_ls.append(filename)
#print "{}->{}".format(filename,dt)
# filename of archive: get the first file, drop the .json extension, and remove all numbers, add the datepattern
file_ls=sorted(file_ls)
stem=re.sub('--*','-', re.sub( '[0-9]','', re.sub('.json$','',file_ls[0]) ))
zipfilename=re.sub('-\.','.', '{}-{}.zip'.format(datepattern,stem))
#print "Zipping up all {} files to {}".format(datepattern,zipfilename)
zfile=zipfile.ZipFile(zipfilename,"w")
for fn in file_ls:
#print "Adding to zip: {} ".format(fn)
zfile.write(fn)
#print "Deleting file: {} ".format(fn)
os.remove(fn)
zfile.close()Note: if you have a backlog of multiple days, you have to run the script multiple times!